Gaussian Processes for Molecular Modeling
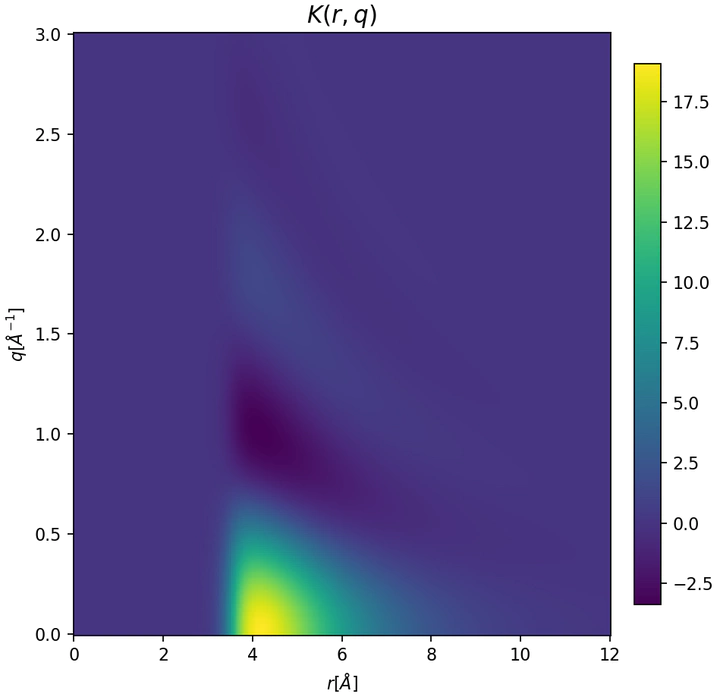
Gaussian processes can model complex experimental data and interatomic interactions while enabling rigorous uncertainty quantification and propagation within a Bayesian framework. We apply state-of-the-art GP methods, including local [1], spectral, and non-stationary kernel design [2], to develop next-generation molecular simulation tools.
[1] B.L. Shanks, et al. Accelerated Bayesian Inference for Molecular Simulations using Local Gaussian Process Surrogate Models, J. Chem. Theory Comput. (2024), https://doi.org/10.1021/acs.jctc.3c01358
[2] Sullivan, H. W., et al. Physics-Informed Gaussian Process Inference of Liquid Structure from Scattering Data, J. Phys. Chem. B. (2025), https://pubs.acs.org/doi/10.1021/acs.jpcb.5c05024