Force Field Design with Bayesian Learning
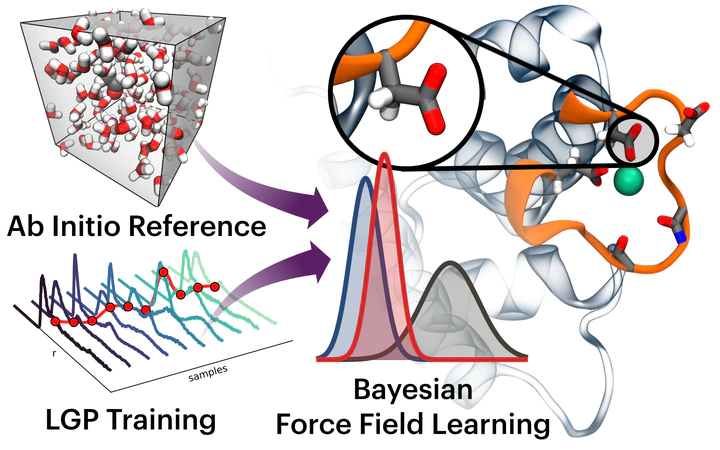
Bayesic Force Fields (BFF) is an open source, Bayesian force field learning tool aimed at addressing transferability and robustness issues with existing biomolecular force fields. The code performs efficient Bayesian inference with physically motivated priors on Coulombic, bonded, and non-bonded terms. Computational feasibility is achieved with a local Gaussian process surrogate model for the classical MD step [1], while large scale ab initio molecular dynamics of the target system is run in parellel to build training observables with quantum level accuracy. The methodology has been used for simple liquids [2], biologically relevant ions in water [3], and charged molecular fragments of proteins, membrane lipids, and carbohydrates [4].
[1] B.L. Shanks, et al. Accelerated Bayesian Inference for Molecular Simulations using Local Gaussian Process Surrogate Models, J. Chem. Theory Comput. (2024), https://doi.org/10.1021/acs.jctc.3c01358
[2] B.L. Shanks, et al. Bayesian Analysis Reveals the Key to Extracting Pair Potentials from Neutron Scattering Data. J. Phys. Chem. Lett. (2024), https://pubs.acs.org/doi/abs/10.1021/acs.jpclett.4c02941#
[3] Fan, S. et al. Charge Scaling Force Field for Biologically Relevant Ions Utilizing a Global Optimization Method. J. Chem. Theory Comput. (2025), https://pubs.acs.org/doi/full/10.1021/acs.jctc.5c00873
[4] Kostal, V. et al. Bayesian Learning for Accurate and Robust Biomolecular Force Fields. J. Chem. Theory Comput. (2026), https://pubs.acs.org/doi/abs/10.1021/acs.jctc.5c02051